Homology Groups - Use as Invariants
As introduced on page here: In two dimensions it is relatively easy to determine if two spaces are topologically equivalent (homeomorphic). We can check if they:
- are connected in the same way.
- have the same number of holes.
However, when we scale up to higher dimensions this does not work and it can become impossible to determine homeomorphism. There are methods which will, at least, allow us to prove more formally when topological objects are not homeomorphic.
These methods use 'invariants': properties of topological objects which do not change when going through a homeomorphism. Here we look at invariants which arise from homology. The page here discusses different invariants which arise from homotopy, which can be harder to work with but may be more selective.
Computing Homology Groups
| When we looked at the delta complex we got a chain of 'face maps' between each dimension and the next lower one. |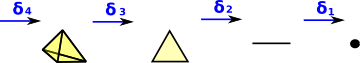  |
In homology we treat this as a chain of Abelian groups.
The boundary of a simplex is not a simplex, but a chain with coefficients 1 or −1. So chains are the closure of simplices under the boundary operator
Here is an outline of an algorithm to compute the homology group from a simplicial complex:
Start with Simplicial ComplexStart with a simplicial complex, in this case 3 vertices with edges forming a circle. |
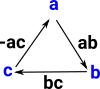 |
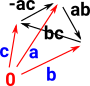
Compute Geometric RealisationConvert vertices, edges and higher order faces to (basis) vectors. So each of the vertices is a vector in its own dimension. They form a set of basis vectors. More about geometric realisation on page here. |
 |
| All the higher order faces are also represented as a set of (basis) vectors. | 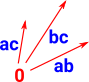 |
Compute Chain ComplexRelate the vertex vectors, edge vectors ... by matrices. More about chain complex on page here. |
Compute Smith Normal FormConvert these matrices to Smith normal form as described below. |
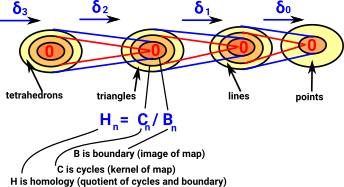
Compute Homology GroupsHomology is cycles modulo boundary. The homology at each step in the chain measures the connectivity of the space. The outer yellow area shows new elements that have been added in at that layer, inside that are elements that have been inherited from higher layers. |
 |
Torsion
The chain complex uses matrices which must be defined over a type which has addition and multiplication. So it can be defined over a Field or a Ring.
If it is defined over a ring of integers (Z) then we have types like Z/2, for example,
Ck=Z Z/2
Z/2
Fractional integer types such as Z/2 are required because the terms in a ring cant be fractional. If the matrices are defined over a field then these torsion types are not required.
Smith Normal Form
The following operations on a matrix are reversible:
- Swapping rows.
- Adding a multiple of one row to another.
- Multiplying a row by an integer.
We can do each of these operations, or combinations of them, by multiplying with a square matrix. Then we can get back to the original matrix by multiplying by the inverse square matrix. We can do the same operations on columns by post-multiplying by the square matrix instead of pre-multiplying.
Doing these operations allows us to get to a diagonal matrix from any arbitrary matrix.
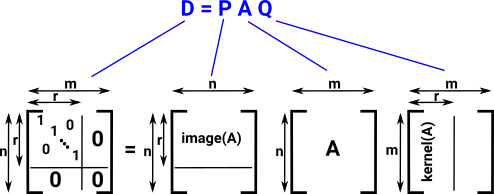 |
Idea for this diagram comes from this website.
The above applies to a matrix over a field or. more specifically over a principle idea domain (PID)
If we work with an integer matrix then, instead of the diagonal of D containing 1's and 0's it now contains invariant factors of A: b1 ≤ b2 ≤...≤ br if bi ≠1 this is torsion Z/a |
|
||||||||||||||||||||||
We use the Smith normal form by calculating from the face maps in linear algebra (matrix) form.
Once in this form we interpret it as follows:
H = Z b1 Z b2
Z b2 Z b3
Z b3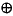 …
…
Homology Groups in FriCAS
|
If A is matrix for Dk+1 and B is matrix for Dk, so that B*A = 0, then torsion part of homology can be read directly from the Smith form of A: rows of leftEqMat give you generators and diagonal entries give order. If diagonal entry is 1, then coresponding cyclic group is trivial and generator is not needed at all. In the case above we get:
as generators of torsion part. Note that torsion part depends only on A, as long as B*A = 0 it does not depend on B. Since A is of full rank in this case free part of homology group is trivial, so the generators above give full description of homology. In general free part of homology may be computed over rational. We can use the following trick: let W be kernel of B treated as matrix over rationals and let V be ortogonal complement of image of A (again over rationals). Now W/Im(A) isomorphic to W \cap V. V is kernel of transpose(A), so to compute W \cap V it is enough to compute kernel of C := horizConcat(B, transpose(A)). You can use completeSmith to get integer basis of this kernel. From Waldek Hebisch post here: |
Homology - Comparison with Homotopy
In homology we don't just use n-spheres but every closed oriented n-dimensional sub-manifold. It also uses a different definition of equivalence classes where composition of loops commutes. This results in Abelian groups. |
 |
So we don't need to fix the basepoint.
If you have not already done so I suggest you read the page about Simplicial Complexes first and also general introduction here.
Homology is an equivalence between cycles in a topological space. On this page we look at code to test this equivalence.
Calculating Homology from Simplexes
Matrix Form of Face Maps
We can represent these face maps by matrix representations, this allows us to relate to linear algebra methods.
We can then generate the homology by putting them in Smith normal form (which is already implemented in Axiom/FriCAS).
Example for Tetrahedron
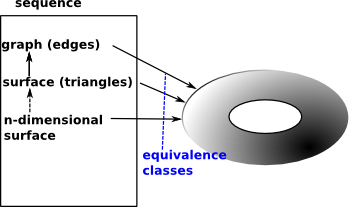
| Here are the face maps as discussed on the delta complex page here. | 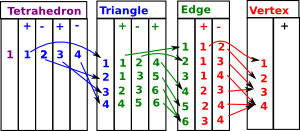 |
We can convert this to a matrices as follows:
| Face map | Matrix | ||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| δ3 = tetrahedron to triangles |
|
||||||||||||||||||||||||||||
| δ2 = triangles to edges |
|
||||||||||||||||||||||||||||
| δ1 = edges to vertices |
|
For these face maps to be valid chain they must be:
- Composible - number of columns in δ1must equal number of rows in δ2.
- Product is zero - δ1* δ2 = 0
Chains
In the case of edges we can chain them together.
A cycle is any combination of edges where at least one decomposition into cycles exists.
Graph Example
H1(X) = Ze-v+1
Orientation and Notation
Getting a consistent orientation naming/numbering is very important. We therefore need to take some time to define it precisely
Lets start with a 2-dimensional face, that is, a triangle: Here it is denoted [a,b,c]. It is oriented, that is, as it is drawn here it has a clockwise orientation. An anti-clockwise triangle such as [a,c,b] can be thought of as an inverse. Here we will denote the edges as bc, ca and ab but it is easier if the nodes are alphabetically ordered, so instead we denote the edges: bc, -ac and ab So now each edge is designated in alphabetical order and the middle edge is negative. |
 |
We can subdivide the triangle by adding another node 'x' as shown in the diagram: This allows us to create 3 triangles. Provided that each of these has a clockwise orientation then, The external edges will have the same direction as the original triangle. The internal edges will each have two edges in opposite directions, so we can think of them as canceling out, leaving us with the original triangle. So, [a,b,c] = [b,c] - [a,c] + [a,b] Designating the triangles in terms of their edges we have: [bc, -ac , ab ] = |
 |
In theory we could divide each triangle up into 4, by splitting each edge, like this: However we don't tend to use this method because:
|
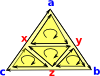 |
Calculated Examples
The only way I could think of to try to validate my FriCAS code is to compare it with the same calculations in SAGE. The sage examples are taken from my SAGE worksheet here. Of course, these are simple examples, so even if they agree it does not necessarily mean my code is correct.
Disc (filled in circle) an 2-sphere examples
gives H0=Z, H1= 0, H2 = 0
| SAGE | FriCAS |
|---|---|
S2 = simplicial_complexes.Sphere(2)
S2.homology()
{0: 0, 1: 0, 2: Z}
S3 = simplicial_complexes.Sphere(3)
S3.homology()
{0: 0, 1: 0, 2: 0, 3: Z} |
(1) -> b1 := sphereSolid(2)$SimplicialComplexFactory
(1)
(1,2,3)
Type: FiniteSimplicialComplex(VertexSetAbstract)
(2) -> homology(b1)
(2) [Z,0,0]
Type: List(Homology)
(3) -> b2 := sphereSolid(3)$SimplicialComplexFactory
(3)
(1,2,3,4)
Type: FiniteSimplicialComplex(VertexSetAbstract)
(4) -> homology(b2)
(4) [Z,0,0,0]
Type: List(Homology) |
Sphere and Disc Surface (Boundary) Example
gives H0=Z, H1= Z, Hn = 0 {n>1}
| SAGE | FriCAS |
|---|---|
BS2 = SimplicialComplex([[0,1], [1,2], [0,2]])
BS2.homology()
{0: 0, 1: Z}
|
(5) -> b3 := sphereSurface(2)$SimplicialComplexFactory
(5)
(1,2)
-(1,3)
(2,3)
Type: FiniteSimplicialComplex(VertexSetAbstract)
(6) -> homology(b3)
(6) [Z,Z]
Type: List(Homology)
(7) -> b4 := sphereSurface(3)$SimplicialComplexFactory
(7)
(1,2,3)
-(1,2,4)
(1,3,4)
-(2,3,4)
Type: FiniteSimplicialComplex(VertexSetAbstract)
(8) -> homology(b4)
(8) [Z,0,Z]
Type: List(Homology) |
Torus example
| SAGE | FriCAS |
|---|---|
RP3 = simplicial_complexes.Torus()
RP3.homology()
{0: 0, 1: Z x Z, 2: Z} |
(9) -> b5 := torusSurface()$SimplicialComplexFactory
(9)
(1,2,3)
(2,3,5)
(2,4,5)
(2,4,7)
(1,2,6)
(2,6,7)
(3,4,6)
(3,5,6)
(3,4,7)
(1,3,7)
(1,4,5)
(1,4,6)
(5,6,7)
(1,5,7)
Type: FiniteSimplicialComplex(VertexSetAbstract)
(10) -> homology(b5)
(10) [Z,Z*2,Z]
Type: List(Homology) |
Real projective space example
| SAGE | FriCAS |
|---|---|
RP4 = simplicial_complexes.
RealProjectiveSpace(2)
RP4.homology() |
(11) -> b6 := projectivePlane()$SimplicialComplexFactory
(11)
(1,2,3)
(1,3,4)
(1,2,6)
(1,5,6)
(1,4,5)
(2,3,5)
(2,4,5)
(2,4,6)
(3,4,6)
(3,5,6)
Type: FiniteSimplicialComplex(VertexSetAbstract)
(12) -> homology(b6)
(12) [Z,C2,0]
Type: List(Homology) |
Klein bottle example
| SAGE | FriCAS |
|---|---|
RP5 = simplicial_complexes.
KleinBottle()
RP5.homology() |
(13) -> b7 := kleinBottle()$SimplicialComplexFactory
(13)
(3,4,8)
(2,3,4)
(2,4,6)
(2,6,8)
(2,5,8)
(3,5,7)
(2,3,7)
(1,2,7)
(1,2,5)
(1,3,5)
(4,5,8)
(4,5,7)
(4,6,7)
(1,6,7)
(1,3,6)
(3,6,8)
Type: FiniteSimplicialComplex(VertexSetAbstract)
(14) -> homology(b7)
(14) [Z,Z+C2,0]
Type: List(Homology) |


